It can be useful to visually indicate the continuation of specific records in a swimmer plot. Optionally adding arrows to the tail ends of swimmer plot lanes in ggswim can help communicate subject survival status, such as whether or not a subject is still on a given study.
To facilitate the addition of arrows, we provide the
geom_swim_arrow() function as a way to tack on these continuation
indicators to your swimmer plot lanes. Behind the scenes, the inclusion
of arrows is facilitated by a call to
ggplot2::geom_segment(), setting zero-length segments so
that assigned arrows are always placed on the right side of indicated
lanes. geom_swim_arrow() gives users control over arrow
neck and head length, along with options for color, fill, and type
(refer to ?geom_swim_arrow for detailed information).
Arrow appearance can be specified in two ways:
- directly via parameters such as
arrow_colour,arrow_fill, andarrow_type - by mapping the arrow aesthetic and supplying styles through
scale_arrow_discrete()
geom_swim_arrow() does not replace the ability to
separately call arrows using the arrow parameter in
geom_swim_lane(). As mentioned,
geom_swim_lane() comes with nearly all of the same
capabilities as geom_segment(), but it may be more
challenging to apply arrows as expected depending on how your data is
structured.
Adding arrows using geom_swim_arrow()
To demonstrate how you might add arrows onto the
patient_data dataset, let’s take a subset of
patient_data that would help us make use of
geom_swim_arrow():
library(ggswim)
library(ggplot2)
arrow_data <- patient_data |>
dplyr::left_join(
end_study_events |>
dplyr::select(pt_id, label),
by = "pt_id"
) |>
dplyr::select(pt_id, end_time, label) |>
dplyr::filter(.by = pt_id, end_time == max(end_time)) |>
dplyr::filter(is.na(label)) |>
unique()
arrow_data
#> # A tibble: 8 × 3
#> pt_id end_time label
#> <chr> <dbl> <chr>
#> 1 04 9 NA
#> 2 09 12 NA
#> 3 13 2.5 NA
#> 4 14 0.9 NA
#> 5 15 0.9 NA
#> 6 17 2.8 NA
#> 7 18 3.3 NA
#> 8 19 6 NAThis should look familiar as a pared-down subset of
end_study_events without an indicated label.
Since filled out label statuses from end_study_events
dataset mean a subject went off study, arrows are only applicable for
subjects with no end study status. Now let’s use
geom_swim_arrow() in combination with
geom_swim_lane() to make a swimmer plot:
patient_data |>
ggplot() +
geom_swim_lane(
mapping = aes(
x = start_time, xend = end_time, y = pt_id,
color = disease_assessment
),
linewidth = 5
) +
geom_swim_arrow(
data = arrow_data,
mapping = aes(xend = end_time, y = pt_id),
arrow_neck_length = 5,
arrow_head_length = grid::unit(0.15, "inches"),
arrow_colour = "firebrick",
arrow_fill = "gold"
)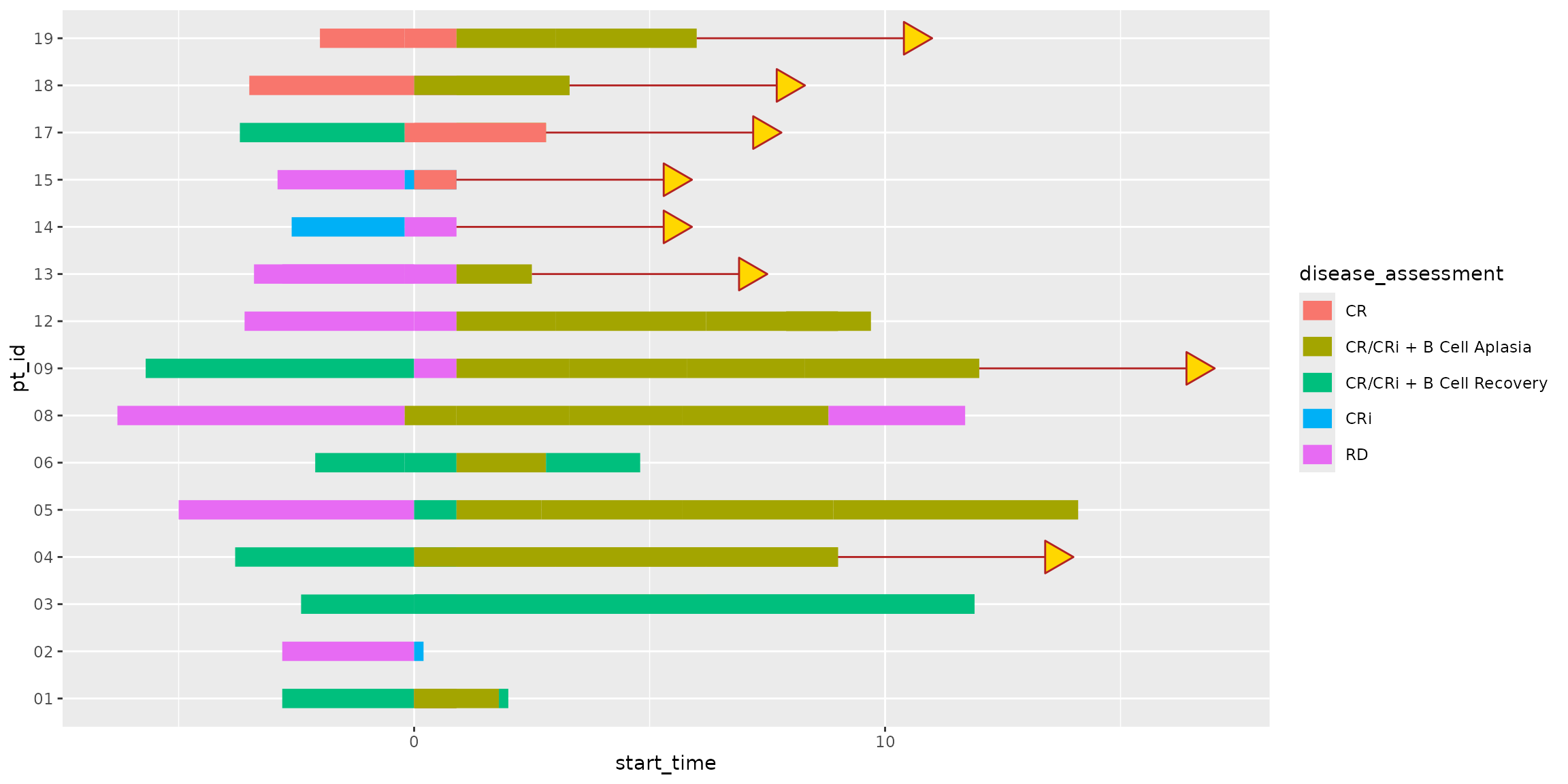
Here we’ve correctly assigned arrows to only the subset of patients
that have not met an end of study event. Note that
geom_swim_arrow() gives plenty of control to the arrow head
and neck color, shape, and even length.
geom_swim_arrow() also allows for aesthetically mapped x
“start” and xend values. If you would prefer to have
dynamically mapped arrow neck lengths indicative of your data, simply
supply the x and xend values in
aes() for geom_swim_arrow() to use.
Using scale_arrow_discrete() for arrow styling
In addition to specifying arrow appearance directly through
parameters, arrows can also be styled using the arrow aesthetic together
with scale_arrow_discrete(). This approach allows arrow
styles to be controlled through the ggplot2 scaling system, making it
easier to add arrows as their own legend component or to manage multiple
arrow styles.
To demonstrate this approach, we can map a value to the arrow
aesthetic and then define its appearance using
scale_arrow_discrete():
patient_data |>
ggplot() +
geom_swim_lane(
mapping = aes(
x = start_time, xend = end_time, y = pt_id,
color = disease_assessment
),
linewidth = 5
) +
geom_swim_arrow(
data = arrow_data,
mapping = aes(
xend = end_time,
y = pt_id,
arrow = "Continuation"
),
arrow_neck_length = 5,
arrow_head_length = grid::unit(0.15, "inches")
) +
scale_arrow_discrete(
limits = "Continuation",
colours = "firebrick",
fills = "gold",
types = "closed",
name = NULL
)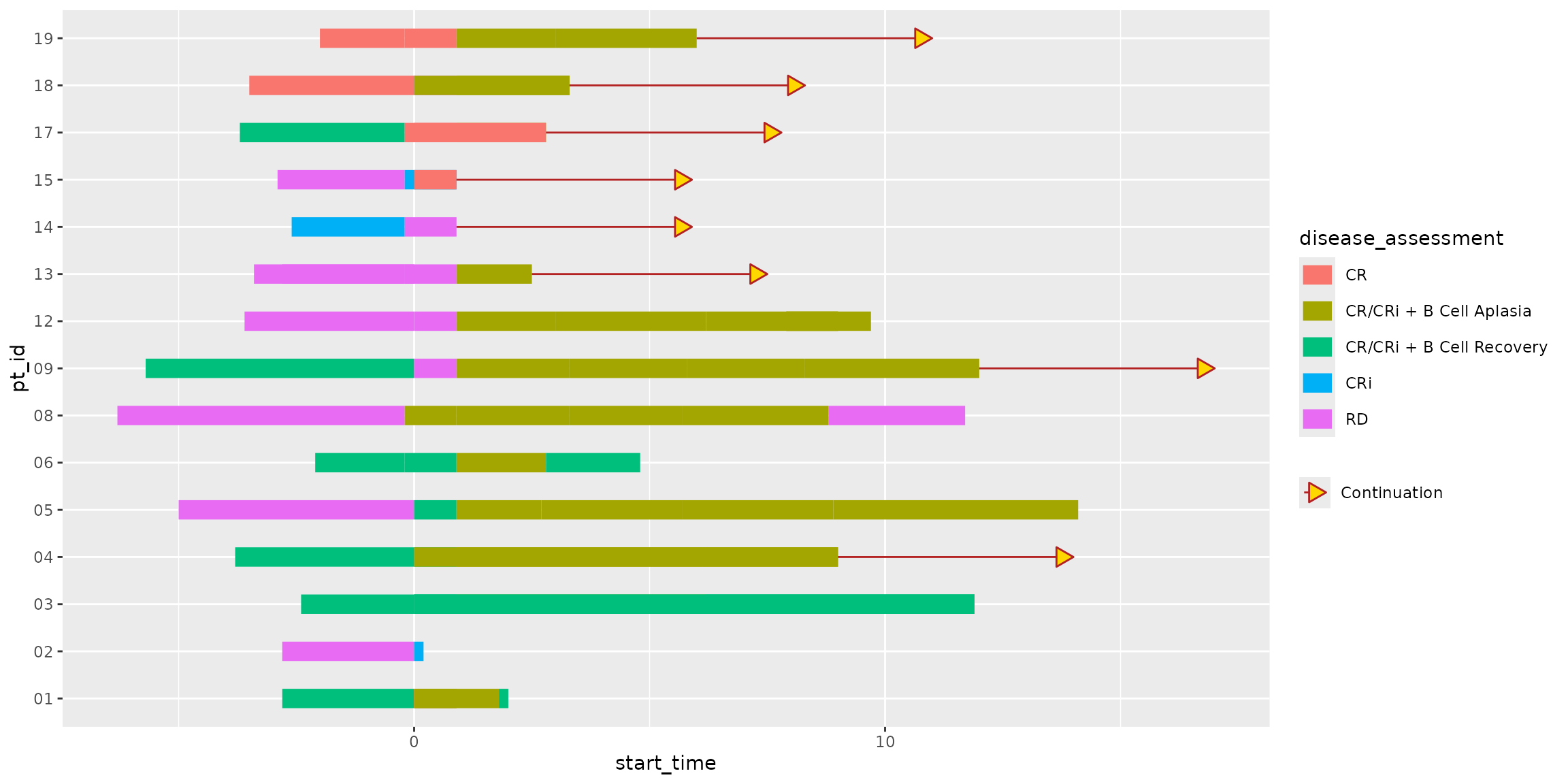
Here, the “Continuation” value mapped to the arrow aesthetic is used
by scale_arrow_discrete() to determine the arrow’s visual
properties. Because this mapping is treated like any other discrete
scale in ggplot2, it can also be displayed as a separate legend entry if
desired.
Using scale_arrow_discrete() is particularly helpful
when:
- arrows should appear as a dedicated legend component
- multiple arrow styles are required
- arrow styling should be managed consistently across plots
For simple cases, specifying arrow parameters directly in
geom_swim_arrow() may still be sufficient. However, using
scale_arrow_discrete() provides a more flexible and
scalable approach when working with more complex swimmer plots.
